The sifter#
This tutorial shows how to use the sifter() function to apply SiFT with a single function call.
We use the Drosophila wing disc development single cell data [1] for the demonstration.
[1]:
import scanpy as sc
import sift
Import data#
download pre-processed data drosopila.annotated.h5ad
[2]:
DATA_DIR = ".../drosophila.annotated.h5ad" # set path to anndata
[3]:
adata = sc.read(DATA_DIR)
Run sifter#
We apply filtering using a mapping kernel (metric="mapping") defined with respect to cell cycle and sex labels (kernel_key="phase_sex") and filtering the gene expression (embedding_key="X").
Additional parameters we can specify are:
copy: bool stating whether theadatais modified in place or copied.pseudocount: add apseudocountto the filtered object to ensure non-negativity ofX.
[4]:
metric_ = "mapping"
kernel_key_ = "phase_sex"
embedding_key_ = "X"
[5]:
sift.sifter(
adata=adata,
kernel_key=kernel_key_,
metric=metric_,
embedding_key=embedding_key_,
copy=False,
pseudocount=False,
)
INFO sift: initialized a SiFTer with mapping kernel.
INFO sift: Filtering cell-cell similarity kernel using projection on `X`.
INFO sift: The data is `SiFTed`!
The filtered embedding is stored in `adata.X`
Finish
Visualize results#
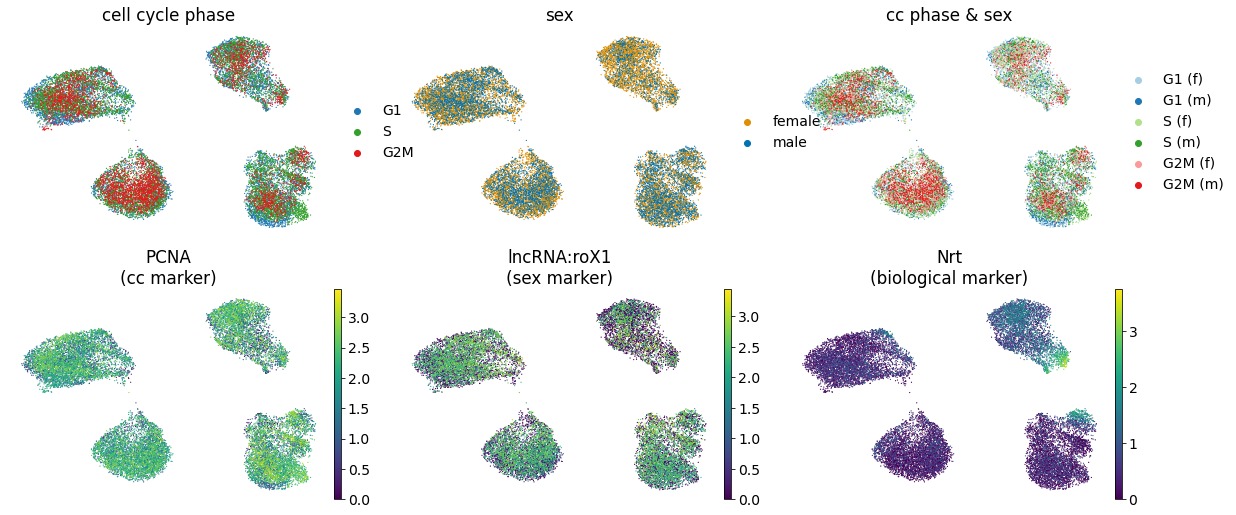
We observe the removal of the nuisance effect with respect to the cell cycle, sex, cc \& sex labels along-with the marker genes PCNA (cell cycle marker), and lncRNA:roX1 (sex marker).
Further, for a biological marker gene, Nrt, we see a pattern is preserved.
[ ]:
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.umap(adata)
[7]:
sc.pl.umap(
adata,
color=["phase", "sex", "phase_sex", "PCNA", "lncRNA:roX1", "Nrt"],
title=[
"cell cycle phase",
"sex",
"cc phase & sex",
"PCNA\n(cc marker)",
"lncRNA:roX1\n(sex marker)",
"Nrt\n(biological marker)",
],
frameon=False,
wspace=0.1,
ncols=3,
)
[ ]:
